FAO/WHO Experts Evaluate Omics Technologies for Microbiological Risk Assessment
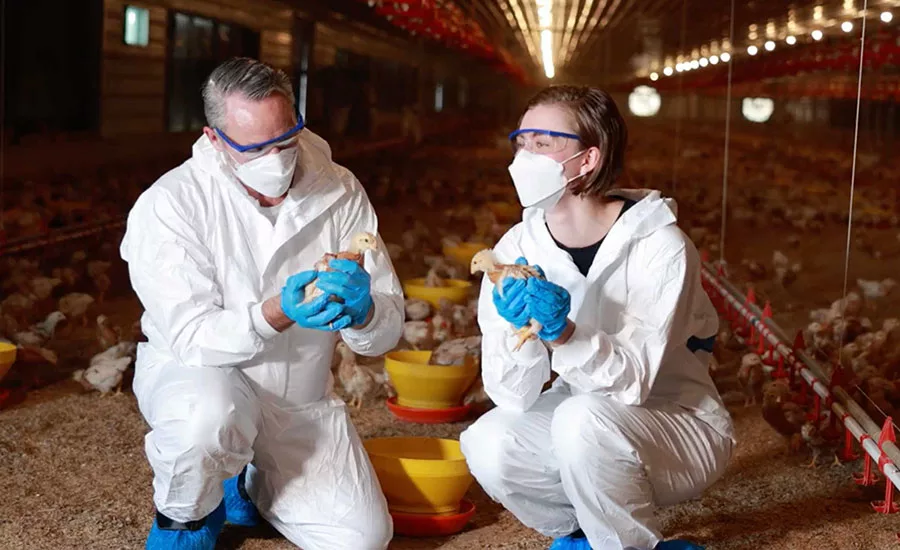
From March 2–6 in Rome, Italy, the Joint Food and Agriculture Organization of the United Nations (FAO)/World Health Organization (WHO) Expert Meetings on Microbiological Risk Assessment (JEMRA) convened to examine how omics-based technologies can be applied to strengthen food safety microbiological risk assessments.
The meeting was convened in response to a request from the Codex Committee on Food Hygiene (CCFH) and focused on whether existing microbiological risk assessment guidance should be updated to reflect scientific advances.
The Omics Landscape in the Context of Microbiological Risk Assessment
The expert committee reviewed the use of genomics, transcriptomics, proteomics, and metabolomics across the four core components of microbiological risk assessments: hazard identification, hazard characterization, exposure assessment, and risk characterization.
Participants concluded that omics-based approaches provide higher-resolution characterization of microorganisms, including their genetic traits, virulence potential, and environmental behavior, compared to traditional classification methods.
Genomics was identified as the most mature and widely applied omics tool in food safety, currently used in surveillance, outbreak investigation, and source attribution.
Other omics approaches, such as proteomics and metabolomics, were described as supporting improved understanding of microbial behavior. However, the committee noted ongoing challenges, including difficulties interpreting genotype–phenotype relationships, distinguishing viable from non-viable organisms, and low pathogen prevalence in foods.
Use Cases for Omics Data
The meeting found that omics data can enhance hazard identification by improving the linkage between specific pathogen subtypes and illness. In hazard characterization, omics data were said to support more refined, subgroup-specific assessments of virulence and dose–response relationships. For exposure assessment, genomics can inform subgroup prevalence and microbial behavior in specific food commodities, although validated genotype–phenotype associations remain limited.
Looking for quick answers on food safety topics?
Try Ask FSM, our new smart AI search tool.
Ask FSM →
Experts also noted that omics-based subgrouping enables more precise risk characterization by integrating differences in virulence, prevalence, and environmental fitness. These capabilities may reduce uncertainty in risk estimates and support more targeted risk management strategies.
Despite the benefits, the committee emphasized that omics data should complement, not replace, traditional microbiological risk assessment approaches. Effective use requires integration with phenotypic, epidemiological, and experimental evidence, along with transparent documentation of analytical methods and assumptions.
Barriers to Adoption
The meeting identified expertise and resource-related barriers to adoption. Capacity-building efforts, particularly in low- and middle-income countries, were recommended to support broader implementation. The committee also highlighted the importance of standardized methods, robust metadata, and adherence to Findable, Accessible, Interoperable, and Reusable (FAIR) data principles.
Potential Codex Guidelines Revisions to Reflect Future Use of Omics
Looking ahead, FAO and WHO experts said that advances such as culture-independent methods, artificial intelligence (AI) analytics, and improved data integration tools are expected to further expand the role of omics in microbiological risk assessment.
The growing challenge of antimicrobial resistance (AMR) and the need for One Health approaches were also identified as key areas where omics technologies can provide value.
The committee reviewed existing Codex Alimentarius guidelines for microbiological risk assessment and management and suggested targeted updates to reflect the emerging role of omics-derived data. A full technical report with detailed recommendations is expected to be published by FAO and WHO.









