The Benefits and Barriers of Whole-Genome Sequencing for Pathogen Source Tracking: A Food Industry Perspective

Much has been written about whole-genome sequencing (WGS) as a tool for outbreak detection and investigation. The point of view from authorities and academia is well-documented, but a broad perspective on its use by the food industry has been missing so far. A workshop was held in September 2019 in Lausanne, Switzerland, bringing together several food companies to define a more general point of view, share experiences around implementation, and explore synergies from working together. Prior to the workshop, a survey was sent out to collect information. Nineteen different companies participated in the survey and/or roundtable discussions. These companies included: Arla Foods, Bonduelle, Cargill, Conagra Brands, Danone, Darling Ingredients, DSM, Fonterra, Greenyard, Hochdorf, Kerry, Kraft Heinz, Mars, Mondelëz International, Nestlé, Unilever, and Vion Food Group. The survey and discussions provided unique insights into how, when, and for what purposes WGS is used by these companies.
WGS and Its Use by the Food Industry
Many food safety authorities around the globe have implemented WGS, or are doing so, for pathogen source tracking. The vision of PulseNet International is for WGS to be used in all public health laboratories to identify, characterize, and subtype foodborne pathogens.[1] Of the 18 companies that participated in the survey, 15 (83%) responded that they have used WGS. However, it should be noted that more companies were invited to participate in the survey/workshop but declined to do so. Therefore, the results described in this article have a positive bias, and it is unlikely that 83 percent of the food industry as a whole has used WGS. The importance of WGS in global foodborne disease surveillance was recognized during the workshop, and the participants recommended that the food industry become familiar with WGS, including the terminology, benefits, and barriers. The level of WGS integration varied between companies, ranging from “not yet used” to “used routinely with in-house resources.” For those companies that have not yet used WGS, the advice would be to have a plan of action on how WGS could be accessed in case an urgent need arises. To help other companies become familiar with WGS, a group of volunteers have taken the initiative to write a guidance document tackling knowledge gaps/hurdles that prevent the use of WGS by the food industry today.
The U.S. Food and Drug Administration (FDA) is sequencing all its historical pathogen isolates spanning 20 years. In addition, all pathogen isolates that are collected through the agency’s current surveillance and inspection programs are being sequenced.[2] There are few companies taking a similar proactive approach of sequencing all their pathogen isolates. Most companies sequence a selection of isolates coming from a particular environmental site or product category, linked to a specific event and after using other subtyping tools. The subtyping tools that were mentioned include serotyping, ribotyping, pulsed-field gel electrophoresis, or matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. WGS is mostly applied when alternative subtyping methods are insufficient to differentiate isolates to a level that will allow companies to define the appropriate action for that particular case. Therefore, it is unlikely that WGS will replace all other typing tools in the near future. Since WGS is not used for all isolates, the definition of “resident” and “transient” can vary from how FDA has defined these terms.[3] Another consideration is the fact that other subtyping tools (e.g., ribotyping) provide insights (e.g., clustering) that may not be compatible with WGS. In this case, it would be important to either build a link between these other tools or sequence all historical isolates.
It was recognized that the proactive sequencing of all pathogen isolates could bring additional insights. However, when weighing the potential benefits against the cost of sequencing and analyzing all pathogen isolates, the balance was not considered favorable. Even more so considering the fact that there are already established systems in place to eliminate pathogens from the value chain and production environment.
Although the level of WGS integration varies between companies, there was clear alignment that WGS has the highest discriminatory power of all subtyping tools and is sometimes the only tool that can give a conclusive answer, laboratory cross-contamination being a case in point. This cross-contamination can originate from proficiency testing scheme (PTS) strains or laboratory control strains or other wild-type isolates being tested in the same facility. For example, if a Salmonella serotype that is commonly isolated from food commodities or factory environments was used in a proficiency test or as a control strain, without WGS, it will not be possible to identify the origin of the strain. It has been proposed to create a common database of these strains to speed up the analysis and reduce duplication of sequencing.
Other Applications of WGS
As described in the literature, WGS data can be used for more than just pathogen source tracking. Examples are predicting adaptation/survival/resistance to biocides, metals, and antimicrobials. It can also be used to predict potential virulence of strains as well as shelf life and spoilage potential of products.[2] Despite this wide range of potential uses, 88 percent of companies participating in this survey question (n = 17) use WGS data mostly for pathogen source tracking, while three companies indicated the use for strain characterization for probiotics (e.g., antimicrobial resistance) and a single company indicated the use of WGS for strain characterization linked to bioproduction or spoilage. Even though the approaches are interesting and can give multiple answers from a single analysis, in some situations, it can be a convoluted approach, whereas a simple phenotypic analysis can provide a quicker answer. Therefore, it is important to match the tool with the question that needs to be answered. Further discussion elucidated that it is unlikely that WGS would be used more if functional characterization becomes more accessible. This requires fundamental knowledge that is difficult to develop in-house and would be easier to outsource.
Barriers to the Use of WGS
A compelling reason for the food industry to embrace a new technology is if its use will lead to operational improvements or accelerate the release decision of products. On one hand, WGS has the highest discriminatory power that can conclusively determine the relatedness of isolates, thereby confirming (or not) the root cause and offering additional insights (e.g., identifying persistent strains). On the other hand, there are significant barriers such as cost, “time-to-results,” complexity of analysis, lack of standardization, and regulatory implications.
Even if the cost of sequencing has been diminishing over the years, WGS is still expensive compared with other subtyping tools.[4] The costs consist of investments in the laboratory and IT infrastructure, equipment, consumables, and the development of expertise in sequencing, bioinformatics, and microbial genomics. Data storage is a concern for the food industry since a powerful IT infrastructure is required, even when WGS analysis is carried out by third parties. An important consideration with regard to cost is not to compare WGS with the standard microbiology detection and identification methods, as it is based on advanced molecular biology technology. A comparison to methods using the same technology such as genetically modified organisms and authenticity testing would put WGS in a more realistic price bracket.
Another barrier is the time-to-results, which is usually more than 1 week. Consequently, product release cannot rely on WGS analysis, and corrective actions need to be taken before. The relative high cost of the analysis that provides a retrospective result can be difficult to justify.
Furthermore, WGS does not give a simple yes/no answer, but it instead provides functions of relatedness of isolates (genetic similarity), which still need appropriate interpretation. Further complexities are linked to the methodology and relevant quality parameters for both the laboratory part (DNA extraction, library preparation, and sequencing), bioinformatics analysis, and, in particular, setting relevant single nucleotide polymorphism/allelic thresholds. Establishing the genetic similarity of isolates requires not only expertise in the laboratory and bioinformatics parts; food safety microbiology expertise is also essential to bring in the wider context of an investigation and will ensure that reliable identification of the root cause and effective corrective actions are defined. It needs to be understood that WGS analysis is only one of the tools available, and other information, especially quality metadata, is essential to get to a meaningful conclusion. Even if WGS analysis is outsourced, internal expertise for all three parts is still needed to correctly interpret the results and relate these to the manufacturing setting.
The lack of method standardization was also highlighted as a barrier. Method standardization through the International Organization for Standardization (ISO) is imperative for accreditation, alignment of methodology, and ensuring results will be internationally recognized. As of today, there is no standard describing the methodology for the use of WGS in pathogen source tracking. In 2014, a new work item proposal was submitted to the general committee for microbiology methods (TC 34/SC 9) and, after acceptance, the working group Whole-Genome Sequencing for Typing and Genomic Characterization was formed to develop a standard for WGS, ISO 23418. A number of members representing the food industry have joined this working group; however, companies that provide sequencing technology or bioinformatics software, and could also provide valuable input in establishing this ISO standard, are currently missing.
PTSs play an important role in the overall acceptance of a methodology. There is a need to have an independent PTS that will be open to all laboratories, including the food industry and third-party laboratories. This will allow comparison of the different methodologies, benchmark the different providers, and assess laboratory proficiency, thereby guiding the choice of laboratory when outsourcing. Such a scheme should cover the entire process, including the laboratory and bioinformatics parts. To our knowledge, the current schemes do not cover both parts and are not open to all laboratories. However, analyzing and interpreting differences in the bioinformatics part will require specific skills that may currently not exist with PTS providers.
Finally, a significant consideration preventing widespread use of WGS by the food industry is the uncertainty around legal/regulatory implications. This included the absence of a legal framework, potential regulatory obligations and/or pressure to share WGS data, and lack of clarity on data ownership (Figure 1). This was highlighted by the survey results and confirmed during the roundtable discussions. Absence of a clear legal/regulatory framework for developing/sharing WGS data and standardized methodology by global regulatory authorities creates uncertainty for companies. There are also concerns about misinterpretation of data generated during internal investigations. It was highlighted that regulators could share examples of cases where companies have been exonerated using WGS, which could help promote the use of and increase confidence in WGS. The decision to use WGS by industry is driven by many factors but should include potential legal risks and regulatory and litigation impacts. It was therefore agreed that the discussions should involve technical and legal experts from the different companies. To kick-start this initiative, it has been proposed to create a common database of laboratory positive control and PTS strains for the benefit of all companies, pending antitrust clearance.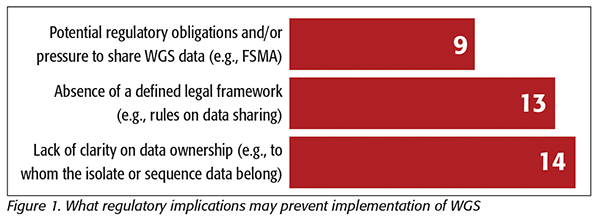
Models of Implementation
Different models of implementation have been adopted by the food industry, ranging from full in-sourcing (i.e., analyzing in-house) to full outsourcing to combinations of the two. The main reasons for in-sourcing are safeguarding confidentiality, ability to influence the time-to-results (shortening it in critical situations), and cost. The last mostly applies to companies with existing sequencing and bioinformatics capabilities that are being used for R&D purposes. To extend these capabilities to pathogen source tracking can be cost-effective, but it may require some modifications.
The laboratory part (DNA extraction, library preparation, and sequencing) is frequently outsourced. Only 2 out of the 17 companies replying to this question have developed in-house capability, and 4 companies use a combination of in- and outsourcing. The main reasons for outsourcing are:
• Capital expenditure investment needed to build in-house capability.
• With the fast-evolving sequencing techniques, sequencers can quickly become obsolete.
• Due to the low sample throughput, the cost per analysis is high with a low return on investment for the equipment.
• DNA library preparation and sequencing are not yet automated.
• Shipping isolates to a central place remains a challenge for some companies.
The bioinformatics part is outsourced less. Here, 5 of the 16 companies replying to this question have developed in-house capability, 5 companies use a combination of in- and outsourcing, and the remaining 6 companies outsource the analysis. The reasons for this are the lack of method standardization and analyzing the data in-house will ensure a consistent approach. It will also facilitate an integrated approach of expertise in genomics and microbiological food safety management expertise.
The selection of third-party laboratories is based on criteria that include acceptable time-to-results, confidentiality, geographic proximity, and competence in genomics. WGS for pathogen source tracking is not like other microbiological methods, such as Salmonella detection. For the food industry to assess the competency of a third-party laboratory in genomics, a specific checklist covering the points that are critical for WGS analysis would be beneficial. Another attention point is around subcontracting to unknown third-party laboratories, which could include authority or governmental laboratories. From discussions, it seems that this type of subcontracting does occur; however, due to data confidentiality, this should be discussed and agreed on with the client beforehand.
Other Genomic Tools and Their Uses
Authorities have started using other genomics tools such as meta-barcoding and metagenomics.[5] Half of the companies responding to the survey indicated that they are also using these other genomics approaches. Of these nine companies, five are using meta-barcoding, two are using metagenomics, and two companies are using a combination. The application of these genomics tools is much broader compared with WGS, as illustrated in Figure 2, which includes the responses of all companies (n = 17 for this question), including those not yet using these tools.
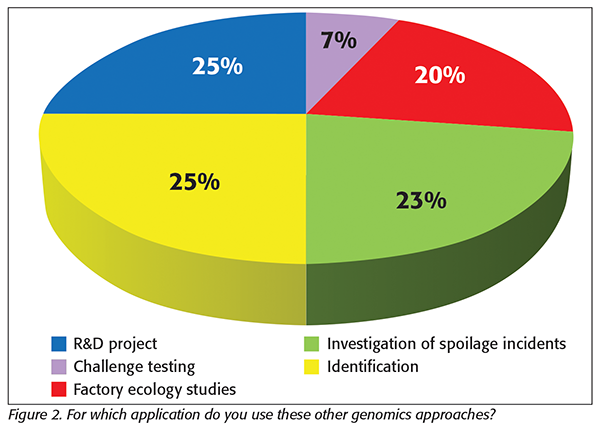
The discussion during the workshop revealed that product development, recipe robustness, and shelf-life studies are examples of R&D projects where these genomics tools are being used. Furthermore, it was highlighted that these tools are particularly interesting for the investigation of spoilage incidents. When comparing spoilage with safety incidents, in the case of the latter, pathogens are normally present at low levels and can be missed by meta-barcoding and metagenomics. Moreover, for these organisms, there are highly specific and sensitive methods that can provide more reliable insights. In spoilage incidents, the organisms are usually present at higher levels, there can be a large diversity, and the methods (e.g., for yeast and molds) are general and will struggle to recover all species, thereby not providing a complete picture. These applications make meta-barcoding and metagenomics interesting new assets in the overall toolbox of food safety testing. In terms of regulatory pressure, it looks like these genomics tools will be easier to implement. However, in terms of practicalities, there are barriers like those for WGS. In particular, development of expertise will represent a considerable investment to build in-house. Building a consortium with several interested food companies could be a more cost-effective approach.
Outlook
The introduction of WGS has been a game changer in the field of analytical microbiology and is expected to be a permanent feature in food safety management systems. The level of WGS implementation by the food industry varies; some members have competencies covering all aspects of WGS analysis, while others are unfamiliar with the tool. Although the advantages of using WGS for public health investigation are evident, the food industry applies WGS mostly when alternative subtyping methods are not sufficient to differentiate isolates. Because the use of WGS by regulatory authorities is likely to intensify, food industry members not yet familiar with it should get on board and become familiar. Understanding the terminology, methodology, and critical points will allow companies to have a discussion on WGS, not only with authorities but also with other food companies and raw material suppliers. The guidance document that is being written will support other food industries in becoming familiar with WGS. To further exchange information among food industries, a network will be established. Please contact FoodMicrobiologyForum@rdls.nestle.com for more information.
Adrianne Klijn leads the Microbial and Molecular Analytics group at Nestlé Research in Switzerland.
Deann Akins-Lewenthal is the senior director of Food Safety and Enterprise Laboratory Services for Conagra Brands in Omaha, NE.
Bala Jagadeesan is senior specialist in the Food Safety Microbiology group at Nestlé Research in Switzerland.
Leen Baert is a specialist in the Food Safety Microbiology group at Nestlé Research in Switzerland.
Anett Winkler is the Europe, Middle East, and Africa regional food microbiologist at Cargill.
Caroline Barretto is a senior bioinformatics specialist at Nestlé Research in Switzerland.
Alejandro Amézquita is a science and technology director within the Foods & Refreshment Division at Unilever.
References
1. Nadon, C, et al. 2017. “PulseNet International: Vision for the Implementation of Whole Genome Sequencing (WGS) for Global Food-Borne Disease Surveillance.” Eurosurveillance 22(23).
2. Allard, M, et al. 2018. “Genomics of Foodborne Pathogens for Microbial Food Safety.” Curr Opin Biotechnol 49:224–229.
3. Pightling, AW, et al. 2018. “Interpreting Whole-Genome Sequence Analyses of Foodborne Bacteria for Regulatory Applications and Outbreak Investigations.” Front Microbiol 9:1482.
4. apps.who.int/iris/handle/10665/272430.
5. Jagadeesan, B, et al. 2019. “The Use of Next Generation Sequencing for Improving Food Safety: Translation into Practice.” Food Microbiol 79:96–115.
Looking for quick answers on food safety topics?
Try Ask FSM, our new smart AI search tool.
Ask FSM →







