Study Links AMR in Salmonella Kentucky to Imported Food
Credit: sbtlneet via Pixabay
A recent study published in Foodborne Pathogens and Disease has provided new insights about the antimicrobial resistance (AMR) of foodborne Salmonella Kentucky. The study’s findings suggest that many human infections of a specific, antimicrobial-resistant sequence type (ST) of S. Kentucky may be linked to the consumption of food products that are imported or consumed while abroad. Additionally, the study found unique differences in the composition of virulence genes and AMR genes among various S. Kentucky clades, which may provide information about the host specificity and pathogenicity of S. Kentucky lineages.
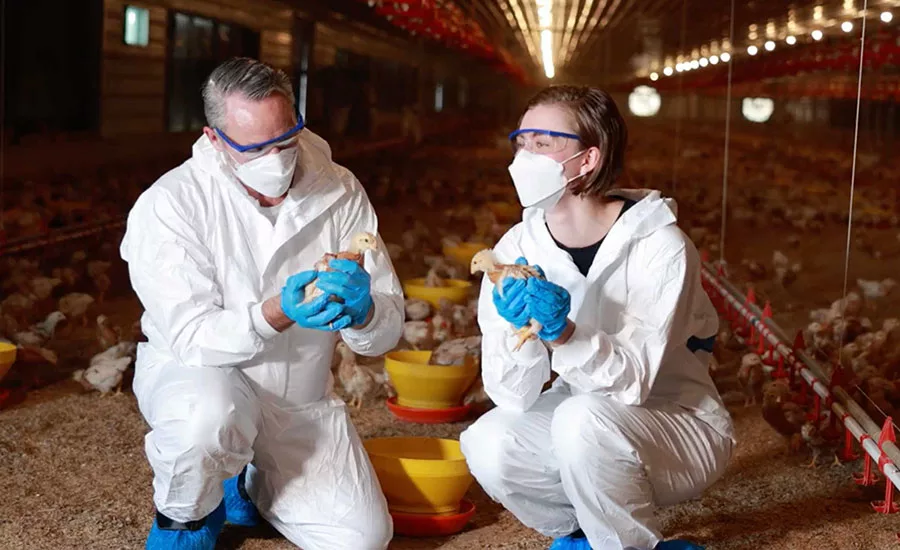
According to data gathered from the U.S. National Antimicrobial Resistance Monitoring System (NARMS), S. Kentucky is frequently isolated from the ceca of chickens (29 percent of isolates) and dairy cattle (3.7 percent of isolates), and is the predominant serovar found in retail chicken products (28 percent of isolates). Despite S. Kentucky’s predominance in certain agricultural animals and agrifood products, historically, the pathogen has caused very few human clinical infections. However, the recent global emergence of multidrug-resistant S. Kentucky strains in humans has created cause for concern, as the trend indicates evolutionary pathways for the serotype toward more virulent subtypes that can become common causes of human illness. Therefore, the study aimed to better describe the genetic diversity, AMR, and virulence characteristics of S. Kentucky isolates sampled from a variety of sources.
Using NARMS data, researchers conducted whole genome sequencing (WGS) analysis on 774 S. Kentucky isolates sampled from humans, food animal ceca, retail meat and poultry products, imported foods and food products, and other samples. Of the S. Kentucky samples that were analyzed, 63 percent of human isolates and 9.2 percent of cecal isolates and retail meat and poultry isolates were ST198. Isolates from imported food belonged to ST198 at a rate of 60 percent. The study’s researchers divided ST198 into distinct clades, one of which comprised almost entirely isolates from humans and imported foods that all contained mutations that lead to fluoroquinolone resistance.
Although human clinical illness associated with S. Kentucky is low, ST198 appears to account for most human infections in the U.S., and the ST is not commonly prevalent among ceca of domestic food animals and retail meat and poultry products. Combined with human exposure data, the findings suggest that fluoroquinolone-resistant ST198 infections may be linked to the consumption of food products that are imported or consumed while abroad. The study also concludes that the presence of virulence genes in some STs and absence in others provides clues to the apparent pathogenicity of S. Kentucky.
Looking for quick answers on food safety topics?
Try Ask FSM, our new smart AI search tool.
Ask FSM →









