Unsung Heroes: State and Local Public Health Officials Innovating in Outbreak Investigations
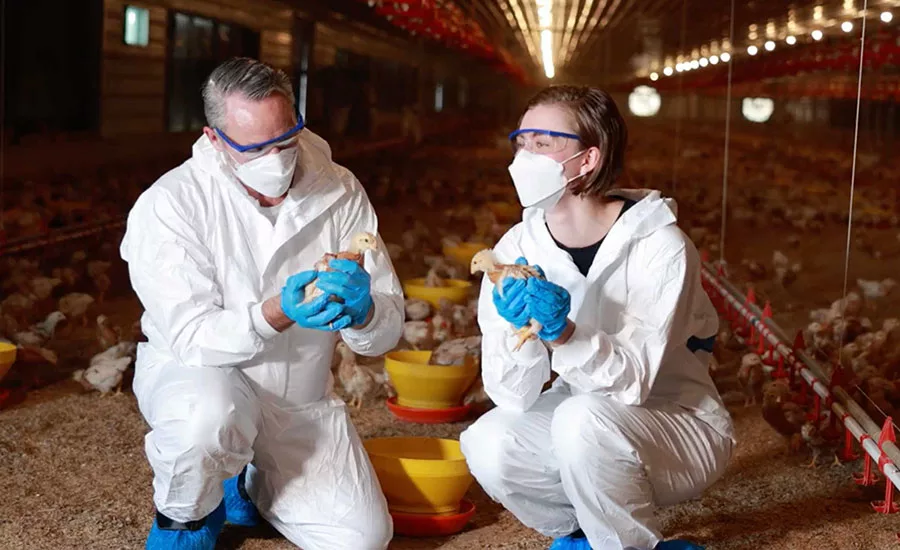
Each year, foodborne illness outbreaks receive extensive media coverage. Typically, the outbreaks that make the headlines are multi-state outbreaks where a group of local, state, and federal public health officials jointly investigate, with federal officials often taking the lead. Each day, state and local officials investigate dozens of intrastate outbreaks with the same level of professionalism and dedication as with a multi-state outbreak. While sometimes these outbreaks are smaller, they still result in significant impact on individuals and communities. This article highlights the work, innovation, and dedication of local and state outbreak investigators in their localized health and agriculture agencies. The innovation and ingenuity at the state and local levels lead to advancements in outbreak investigations on the national level. Even in a multi-state outbreak, these state and local officials are the boots on the ground, providing the information needed to coordinate the national investigation.
An outbreak investigation requires an interdisciplinary approach often referred to as a three-legged stool. Critical for a successful outbreak investigation, the three essential disciplines are environmental health, epidemiology, and laboratory services (see “Essential Disciplines”). Each outbreak investigation is different and may require a different level of effort from each discipline. Successfully investigating outbreaks requires strong programs in each of the three disciplines.
This article will focus on outbreak investigations in three states: Washington, Tennessee, and New York. Each investigation has its own unique nuisances and challenges, but there are themes across the outbreaks, including the use of environmental sampling to determine whether relationships exist between the pathogens found in ill individuals and in the environment of the suspected source. Whole-genome sequencing (WGS) has also become an essential tool in outbreak investigations, assisting in determining how closely genetically related pathogens are. This helps investigators determine the likely source of an outbreak and whether cases appear to be linked. Further, the federal government through the U.S. Centers for Disease Control and Prevention (CDC) and the U.S. Food and Drug Administration (FDA) assist with financial support for state and local outbreak investigations through a variety of funding instruments (see “Programs That Assist State and Local Governments in Outbreak Investigations”).
Washington RRT: Creating Collaborations to Link Illnesses and Adulterated Product across Time
Washington State has developed and continually improved their food and feed Rapid Response Team (RRT) since 2009. As one of the original states to receive FDA cooperative agreement funding, Washington has worked hard to create a flexible, capable response team to effectively mitigate public health threats associated with human and animal food. The Rapid Response Program, housed within the Washington State Department of Agriculture (WSDA), maintains close collaborative relationships with the FDA Office of Human and Animal Foods Division 6-West (OHAF-6W), the Washington State Department of Health (WADOH), local health jurisdictions in Washington State, industry, and other key partners to help safeguard the food and feed supply.
One such capability of the Washington RRT is to quickly coordinate response partners and leverage their resources to link pathogens found in food samples to human illnesses at the genetic level. These illnesses may even be “historical” in nature in that they may have been reported months or years in the past. However, with stronger technologies and methods such as WGS and bioinformatics, these illnesses can now be associated with pathogens collected from current product and clinical samples to a high degree of accuracy. Linking such illnesses to samples collected during current RRT responses may allow responders to identify chronic issues at a food production facility such as a resident pathogen that is not being killed through current cleaning and sanitation procedures.
This type of “rapid response across time” was exemplified through a multi-agency response coordinated by the Washington RRT in September 2017. Eight months earlier, the WSDA Rapid Response Program was notified by foodborne illness epidemiologists at WADOH that two human Salmonella Dublin cases with indistinguishable pulsed-field gel electrophoresis (PFGE) patterns and similar illness onset dates had reported consuming retail raw milk from a specific licensed raw milk dairy in Washington State. With proper licensing through WSDA, retail raw milk and hand-skimmed raw cream are legal in Washington State. Approximately 32 firms were licensed to produce and distribute these raw dairy products within the state’s borders as of December 2018.
Subsequent sampling of the finished raw milk product processed by the dairy in question did not indicate the presence of Salmonella, but rather Shiga toxin-producing Escherichia coli. Based on these lab results, the dairy decided to voluntarily recall the implicated lots of their organic retail raw milk in February 2017.
Fast-forwarding to September 2017, routine surveillance sampling conducted by WSDA subsequently confirmed results for the presence of Salmonella spp. in retail raw milk product processed by the same firm. This time around, the firm respectfully declined to issue a voluntary recall. Due to the possible public health implications based on the positive sample results, WSDA developed and distributed a public health alert, a public information release that includes the sample results, health information on salmonellosis, and distribution information for the positive product.
Through coordination efforts led by the Washington RRT, the WADOH Food Safety Program notified local health jurisdictions throughout the state and provided them with a copy of the alert to be used as a tool for notifying retail points of sale.
After the four additional raw milk samples were confirmed for the presence of Salmonella spp., Washington RRT was notified by WADOH epidemiologists that serology identified the pathogens as Salmonella Dublin. PFGE patterns were also indistinguishable from both the previous product samples and the two human Salmonella Dublin cases back in January 2017. The association between a historical illness cluster and a food product was getting stronger.
With this tie between confirmed positive product samples and historical human illnesses, WADOH issued their own public information in collaboration with WSDA. The release stated that health officials urged consumers not to drink the organic retail raw milk with the implicated sell-by dates from the specific dairy. Shortly thereafter, WSDA issued a summary suspension of the dairy’s milk processing plant license, therefore administratively halting the ability to legally process and distribute retail raw milk. The dairy’s milk producer licenses were not affected by the order, which allowed the dairy to still milk their cows, hold their milk, and divert it to be pasteurized if they chose to do so.
Additionally, Washington RRT and WADOH epidemiologists requested CDC bioinformatics to conduct a genomic comparison of the Dublin isolates. The comparison indicated that all seven samples (five product and two clinical) were highly related to one another within 0–2 single nucleotide polymorphisms.
Leveraging the capabilities of its public health partners and the current gold standard in food safety-related DNA sequencing, Washington State was able to associate human illnesses with a food product and take regulatory steps to mitigate future illnesses. Working closely with the WSDA Food Safety Program, the dairy took specific corrective and preventive measures that hopefully will reduce the risk of product adulteration in the future.
Identifying the capabilities of its response partners and coordinating their deployment to protect public health is a key strength that the Washington RRT brings to an integrated food/feed safety system both within Washington State and nationally. Due to the proactive nature of the public health agencies in Washington State, the newest tools and resources can be effectively utilized to control current public health threats and prevent them from occurring in the future.
Getting to the Problem: Detection of Norovirus in Tennessee
Norovirus is the leading cause of foodborne illnesses in the U.S., with annual illnesses estimated in the millions, thousands of hospitalizations, and hundreds of deaths. The health and economic impacts are significant and keep this pathogen in the forefront of public health efforts. Recent advances in norovirus detection have allowed public health agencies to improve investigation techniques in outbreaks where norovirus is the culprit. Environmental sampling provides a valuable resource in these outbreaks, strengthening the connection between ill people, suspected foods, and environmental contamination.
Norovirus outbreak investigations typically involve three complementary areas: 1) environmental investigations, 2) epidemiological studies, and 3) laboratory analysis of clinical specimens. Sometimes these investigation methods are ineffective in determining etiology and mode of exposure due to people’s unwillingness to provide a sample, small sample size for statistical analyses, and/or poor food history recall. Environmental sampling, however, has helped during outbreak investigations with missing data or complemented outbreaks where data from other areas are strong. Environmental sampling for norovirus is a relatively new approach providing an additional tool to help fill informational gaps and aid in determining the cause of the outbreak, while facilitating short- and long-term control measures.
The Tennessee Department of Health (TNDOH) has worked alongside its public health lab to add this tool to its public health toolbox and has collaborated to develop new sampling protocols that include swabbing for norovirus during outbreaks at foodservice establishments (FSEs). The sampling technique and materials used were based on the environmental sampling protocol described in a recent study.[1] TNDOH has used a swab with a tip made from macrofoam—a material proven to be more effective in recovering norovirus from surfaces when compared with materials like cotton, nylon, and traditional sponges. Additionally, macrofoam has proven effective in releasing norovirus particles during the extraction process.
Since testing this strategy, TNDOH successfully detected norovirus in two restaurant outbreaks. In one of these outbreaks, the local health department was notified that a dining party had become ill with vomiting and diarrhea following meals at a restaurant (Table 1). Illness onset, symptoms, and duration were consistent with norovirus. This prompted an environmental assessment at the restaurant, where a food handler was identified as having diarrhea and being present at the restaurant during the period in question. The employee reported having a diarrheal episode in a single-occupancy, unisex restroom that was not used by any members of the dining party. Investigators used this information to determine likely contamination zones and collected 24 swabs in the ill food handler’s workstation, the restroom used by the ill food handler, the restrooms used by the dining party, and the private dining area (Table 2).

Epidemiological information for this outbreak was limited to the dining party. No additional ill or well customers outside the dining party were identified because customer contact information was not retained by the FSE. Therefore, definitive epidemiological studies could not be performed.
Stool specimens were collected from four out of the eight dining party members who reported illness. TNDOH could not obtain stool from the ill food handler. All stool specimens were positive for norovirus GII.2. Of the 24 swabs collected at the restaurant, two were positive for norovirus GII.2, the same genotype the ill customers had. Importantly, both positive samples were recovered from the same toilet area used by the ill chef who reported having diarrhea while working. All samples recovered from the restroom used exclusively by the dining party were negative.
An environmental assessment revealed that reporting and exclusion policies for ill workers needed to be revised. While the restaurant had an ill-worker reporting policy, employees were not well-trained and were not following the policy. Results of the assessment were communicated to the restaurant, which facilitated better understanding of the importance of training and communication of these food worker policies. Furthermore, environmental sample results directed more intensive cleaning and disinfection of contaminated areas.
Environmental testing for norovirus is a valuable tool to support epidemiological and/or environmental data while providing investigators with an improved understanding of how and why the outbreak occurred. In addition, the environmental sampling results have helped focus remediation efforts by the restaurant. Management was informed of specific areas most likely contaminated so that more aggressive disinfection methods could be deployed. These control measures would not have been recognized and emphasized without the evidence provided from environmental sampling. Adding environmental testing for norovirus to the public health toolbox enhances the ability to understand and respond to outbreaks.
Cracking the Case: Listeria monocytogenes in New York
Listeria monocytogenes is one of the leading causes of fatal foodborne infections in the United States among patients with laboratory-confirmed infections.[2] Reported outbreaks of Listeria infections in the 1990s were primarily linked to deli meats and hot dogs. Today, Listeria outbreaks are often linked to dairy products and produce.[3] Compared with other common foodborne pathogens, L. monocytogenes has a longer incubation period—typically between 14 and 21 days but can be as long as 70 days.[4] The relatively long incubation period of L. monocytogenes makes it difficult to identify the implicated food item that caused the illness. To improve the identification of the source of illness associated with L. monocytogenes cases and outbreaks, the New York State Department of Health (NYSDOH) has been actively implementing the use of WGS since 2013 and of environmental sampling as techniques to solve L. monocytogenes outbreaks.
In 2015, two L. monocytogenes cases were reported to NYSDOH from the same county. Both cases were male, both were in their 70s, one case was hospitalized, and one case was deceased. Both cases were PFGE matches to each other and to other cases nationwide. WGS analysis identified that these two cases were 0–1 alleles apart from each other, but far away from other PFGE matching cases, suggesting a high genetic relatedness. One case and one proxy were reinterviewed with the Listeria Initiative Case Report Form, an enhanced surveillance form designed to collect detailed information on listeriosis cases in the United States. Review of the questionnaires indicated both cases shopped at the same location of Grocery Store A, which has a shopper card program. Purchase histories for both cases were obtained and review of purchase histories indicated that both cases purchased Brand B cheese and deli items from the same deli department within 3 days of each other.
Grocery stores in New York State are regulated by the New York State Department of Agriculture and Markets (NYSDAM). NYSDOH asked NYSDAM to conduct a site visit at Grocery Store A to collect environmental samples at the deli. Twenty environmental samples were collected and submitted to the NYSDAM Food Laboratory for L. monocytogenes detection and PFGE. Subsequently, 11 of 20 environmental samples from the deli were positive for L. monocytogenes. Among these samples, three different PFGE patterns were identified, and 8 of 11 samples were a PFGE match to the outbreak strain. WGS analyses conducted by NYSDOH’s Wadsworth Center identified an environmental sample collected from a cheese tray that was 0–1 alleles apart from the clinical cases, indicating it was highly related and the likely source of illness.
In 2014, a cluster of three L. monocytogenes cases with a matching PFGE pattern were identified in County C. Limited information was available for two of three cases; no epidemiological link was recognized at the time and WGS was not routinely conducted then. Late in 2016, County C reported two additional PFGE matching cases; WGS results indicated these two cases were 0–2 alleles apart and highly related to each other. The 2016 cases both reported consuming food from Establishment D. County C initiated an investigation at Establishment D. In January 2017, an additional case with a PFGE pattern matching the 2014 and 2016 cases was identified, and this case also reported consuming food from Establishment D. WGS analyses conducted by the Wadsworth Center indicated that all six cases from 2014, 2016, and 2017 were highly related to one another (exhibiting a 1–6-allele difference). Additional interviews were conducted, with five of six cases reporting consuming food from Establishment D. County C conducted a site visit at Establishment D and collected 20 environmental samples; however, all 20 samples were reported as negative for L. monocytogenes by the Wadsworth Center. Per recommendation of the county, Establishment D cleaned and sanitized their entire facility in February 2017.
In May 2017, County C reported another PFGE matching case who consumed food from Establishment D. WGS analysis indicated that this case was highly related to the six previous cases. County C collaborated with the NYSDOH Environmental Health Specialist Network (EHS-Net) program, the New York Integrated Food Safety Center of Excellence, and NYSDAM to develop a new environmental sampling plan. Environmental sampling supplies were provided by the NYSDOH EHS-Net program, and environmental sampling was repeated with assistance from NYSDAM; 42 environmental samples of the food production facility and equipment were collected, and 4 of the samples were positive for L. monocytogenes. The four environmental samples were a PFGE match to one another and to the seven clinical cases. WGS analysis indicated that these samples were highly related to the clinical cases. The four samples were collected from a bread rack dolly, a swinging door, and two different floor mats. It was recommended that Establishment D hire a consultant, replace the old floor mats, and clean and sanitize the entire facility. They were forbidden from serving ready-to-eat foods until cleaning and sanitizing was completed and repeat environmental sampling indicated L. monocytogenes was no longer present in the facility.
The use of WGS and collaborative environmental sampling by different agencies to solve L. monocytogenes outbreaks is a proven success story. Epidemiology, environmental health, or laboratory services individually cannot solve an outbreak. The three-legged stool approach combining epidemiological information collected from patients, environmental health information collected from establishments, and WGS results from the laboratory allow us to detect more clusters of illness, link more cases to a likely source, identify unrecognized sources of illness, and stop outbreaks before they escalate in size and severity.
Conclusions
These three foodborne illness outbreak investigations exemplify the dedicated professionals working throughout the United States. Their efforts result in reducing the number of potential illnesses and finding root causes that can assist in prevention of similar illnesses in the future. While typically behind the scenes, professionals managing the outbreaks at state and local health departments are often the unsung heroes of public health.
Disclaimer: Development of the TNDOH section was provided by funding through CDC’s EHS-Net Food grant.
Randy J. Treadwell, M.P.H., is program manager, rapid response & emergency management at the Washington State Department of Agriculture. D. J. Irving, M.P.H., REHS, is an environmental health specialist at the Tennessee Department of Health. David C. Nicholas, M.P.H., is chief epidemiologist at the New York State Department of Health. Steven Mandernach, J.D., is the executive director of the Association of Food & Drug Officials and a member of the Editorial Advisory Board of Food Safety Magazine.
References
1. Park, GW, et al. 2015. Appl Environ Microbiol 81(17):5987–5992.
2. Barton Behravesh, C, et al. 2011. J Infect Dis 204:263–267
3. www.cdc.gov/listeria/prevention.html.
4. Heymann, DL. Control of Communicable Disease Manual, 20th ed. (American Public Health Association, 2015).
Looking for quick answers on food safety topics?
Try Ask FSM, our new smart AI search tool.
Ask FSM →







