Analysis of Pesticides in Baby Foods Using Triple Quad GC/MS/MS
Contamination of food products with pesticides is a growing concern because of recognized adverse health effects, increasing worldwide usage of pesticides, and increasing imports of raw foodstuffs from foreign sources. The concern is particularly acute for baby foods because of the high vulnerability of babies to health effects from synthetic chemicals such as pesticides.
Gas chromatography/mass spectrometry (GC/MS) has been used extensively to identify and quantify trace-level pesticides in food matrices; the most significant challenges have been matrix interference and achievement of meaningful health-based detection limits for the compounds of interest. The QuEChERS (Quick, Easy, Cheap, Effective, Rugged and Safe) sample preparation method[1] has helped to overcome some of the problems of matrix interference, and commercialization of QuEChERS kits has promoted widespread screening of foodstuffs for trace pesticides. But matrix interferences still present a significant challenge for analysis of trace-level pesticides in foods, even after QuEChERS extraction and cleanup.
Triple quadrupole GC/MS/MS has emerged as an important technique for analysis of trace-level contaminants in complex matrices. Operation of the triple quadrupole GC/MS/MS in the multiple reaction monitoring (MRM) mode provides unmatched sensitivity and selectivity for detection and quantitation of target pesticides at low concentrations in the presence of background interferences. Most co-extracted matrix interferences are minimized or completely eliminated using the MRM mode.
This application note presents instrument configuration, operating parameters, and analytical results for analysis of 36 pesticides from various chemical classes in a QuEChERS extract of baby food using triple quadrupole GC/MS/MS. Results were evaluated for calibration linearity, analytical precision and accuracy in a baby food matrix. Selectivity as a function of variable mass spectral resolution settings on the two sets of quadrupoles, Q1 and Q3, is also discussed.
Experimental
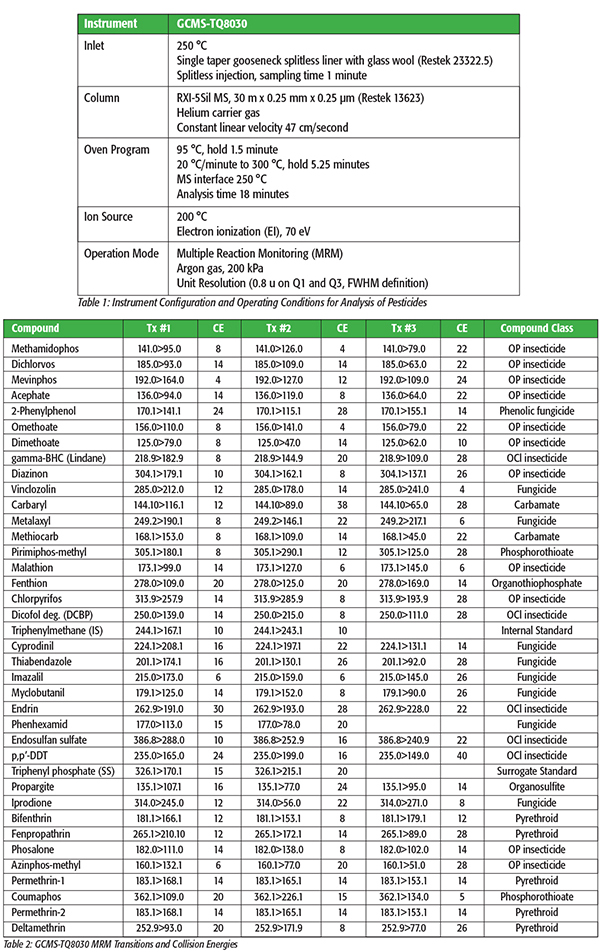 The analyses were conducted using a Shimadzu GCMS-TQ8030 triple quadrupole GC/MS/MS. The GCMS-TQ8030 was operated in the MRM mode, using the optimized MRM transitions and collision energies detailed in the Shimadzu GC/MS/MS Pesticide Database.[2] The GCMS-TQ8030 allows optimization of the collision energy for each MRM transition, providing ultimate sensitivity. The instrument configuration and operating conditions, including MRM transitions and optimized collision energies are shown in Tables 1 and 2.
The analyses were conducted using a Shimadzu GCMS-TQ8030 triple quadrupole GC/MS/MS. The GCMS-TQ8030 was operated in the MRM mode, using the optimized MRM transitions and collision energies detailed in the Shimadzu GC/MS/MS Pesticide Database.[2] The GCMS-TQ8030 allows optimization of the collision energy for each MRM transition, providing ultimate sensitivity. The instrument configuration and operating conditions, including MRM transitions and optimized collision energies are shown in Tables 1 and 2.
A sample of blended pears was used as the test sample matrix; an organic variety was selected so it would be free from background pesticide contamination. The sample matrix was extracted and subjected to cleanup using the QuEChERS procedure. Calibration was conducted using the matrix-matched internal standard procedure. The sample preparation did not involve concentrating or diluting the sample, so concentrations expressed in ng/mL (parts-per-billion, ppb) in the calibration standards and extracts are equivalent to ng/g (parts-per-billion, ppb) in the original sample.
Results and Discussion
Chromatography
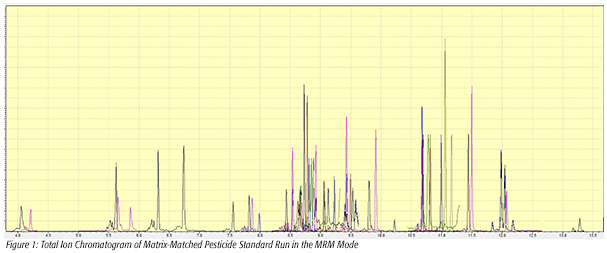 The total ion chromatogram (TIC) acquired in the MRM mode for the pesticide standard is shown in Figure 1, illustrating the chromatographic separation of the target pesticides in this study. In the MRM mode, the TIC for each analyte is the sum of the signal for each MRM transition for that particular compound, so the appearance of the chromatogram is slightly different than a typical TIC acquired in the full-scan mode. The effect of initial column temperature on chromatographic performance is important when the QuEChERS procedure is used; the injection solvent, acetonitrile, produces some unusual chromatographic effects at temperatures below 90 °C.[3]
The total ion chromatogram (TIC) acquired in the MRM mode for the pesticide standard is shown in Figure 1, illustrating the chromatographic separation of the target pesticides in this study. In the MRM mode, the TIC for each analyte is the sum of the signal for each MRM transition for that particular compound, so the appearance of the chromatogram is slightly different than a typical TIC acquired in the full-scan mode. The effect of initial column temperature on chromatographic performance is important when the QuEChERS procedure is used; the injection solvent, acetonitrile, produces some unusual chromatographic effects at temperatures below 90 °C.[3]
Matrix-Matched Calibration
Using the matrix-matched calibration approach, five calibration standards were prepared in a blended pears extract over the range of 1–200 ng/mL (ppb). Triphenylmethane was used as the internal standard and was held constant at 10 ng/mL; triphenyl phosphate was used as a surrogate standard at 20 ng/mL in all standards. The calibration standards were analyzed using the instrument conditions outlined above. The detector voltage (electron multiplier) was adjusted to give acceptable response at the lowest calibration level and avoid saturation at the highest calibration level.
.jpg) Response factors were calculated and percent relative standard deviation (%RSD) determined using GC/MS solution software. The precision of the calibration was evaluated using the correlation coefficient (r) and %RSD of the response factors for each analyte, as tabulated in Table 3. Linearity, as evaluated by the correlation coefficient, was 0.999 or better for all 36 compounds. In many cases, where the RSD for the response factors was greater than 20% (e.g., thiabendazole and imazalil), the presence of native pesticides in the matrix contributed to the signal for the lowest concentration standards and accounts for the high %RSD. When the low-level calibration standard is not included in the calculation, RSD is less than 20% overall.
Response factors were calculated and percent relative standard deviation (%RSD) determined using GC/MS solution software. The precision of the calibration was evaluated using the correlation coefficient (r) and %RSD of the response factors for each analyte, as tabulated in Table 3. Linearity, as evaluated by the correlation coefficient, was 0.999 or better for all 36 compounds. In many cases, where the RSD for the response factors was greater than 20% (e.g., thiabendazole and imazalil), the presence of native pesticides in the matrix contributed to the signal for the lowest concentration standards and accounts for the high %RSD. When the low-level calibration standard is not included in the calculation, RSD is less than 20% overall.
Precision and Accuracy
Eight replicate injections of the 1.0 and 5.0 ng/mL matrix-matched standards were analyzed to assess the precision and accuracy of the method near the low end of the calibration range. The mean concentration and %RSD for the replicate analyses are presented in Table 3.
In conjunction with the precision and accuracy study, a matrix blank (unspiked sample matrix) was analyzed to check for background levels of the target compounds. Discrete chromatographic peaks for approximately one-third of the target compounds were observed in the matrix blank, indicating that these pesticides were present in the native baby food matrix. Concentrations of the native pesticides in the matrix were all below 1.0 ng/mL, and account for recovery anomalies at the lowest spike levels (1.0 ng/mL). The effect of native pesticides in the calibration standards can be avoided by preparing standards in solvent instead of matrix.
Selectivity as a Function of Mass Spectral Resolution
 Resolution settings on the GCMS-TQ8030 are expressed using the FWHM (Full Width at Half Maximum) definition. Three resolution settings are available for each set of quadrupoles when operating in the MRM mode (Table 4), and the resolution can be defined independently for Q1 and Q3, using any combination of the three settings. In most cases, Unit resolution (0.8 microns) for both Q1 and Q3 provides the best combination of sensitivity and selectivity; however, Q1 and Q3 mass spectral resolution settings can be adjusted individually for each analyte to provide a customized method when needed.
Resolution settings on the GCMS-TQ8030 are expressed using the FWHM (Full Width at Half Maximum) definition. Three resolution settings are available for each set of quadrupoles when operating in the MRM mode (Table 4), and the resolution can be defined independently for Q1 and Q3, using any combination of the three settings. In most cases, Unit resolution (0.8 microns) for both Q1 and Q3 provides the best combination of sensitivity and selectivity; however, Q1 and Q3 mass spectral resolution settings can be adjusted individually for each analyte to provide a customized method when needed.
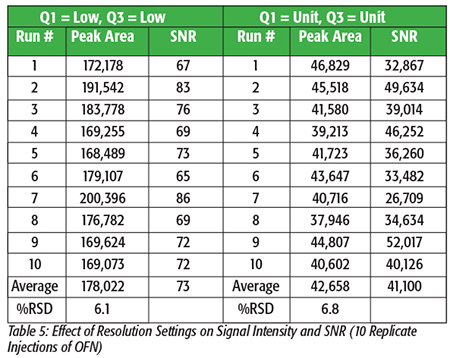 The resolution settings that are chosen can have an enormous impact on signal intensity, noise levels, compound detection limits, and selectivity against background interferences. Low resolution settings (e.g., 3.0 microns) increase analyte signal intensity because more ions strike the detector, but they also increase the corresponding noise levels, so the resulting signal-to-noise ratios (SNR) are actually reduced, and analyte detection limits go up. Unit (0.8 microns) or High (0.6 microns) resolution settings reduce overall signal intensity (fewer ions to the detector), but noise levels are also significantly reduced so SNR and analyte detection limits can be greatly improved. This principle is illustrated using 10 replicate injections of an octafluoronaphthalene (OFN) standard, as shown in Table 5. Even though signal intensity using the Q1=Low and Q3=Low (Low/Low) resolution setting is approximately 4 times higher than when using the Unit/Unit setting, the corresponding noise levels are so high with Low/Low resolution that SNR is dramatically and negatively impacted. In general, the Unit/Unit setting provides the best overall balance of high signal intensity and low noise.
The resolution settings that are chosen can have an enormous impact on signal intensity, noise levels, compound detection limits, and selectivity against background interferences. Low resolution settings (e.g., 3.0 microns) increase analyte signal intensity because more ions strike the detector, but they also increase the corresponding noise levels, so the resulting signal-to-noise ratios (SNR) are actually reduced, and analyte detection limits go up. Unit (0.8 microns) or High (0.6 microns) resolution settings reduce overall signal intensity (fewer ions to the detector), but noise levels are also significantly reduced so SNR and analyte detection limits can be greatly improved. This principle is illustrated using 10 replicate injections of an octafluoronaphthalene (OFN) standard, as shown in Table 5. Even though signal intensity using the Q1=Low and Q3=Low (Low/Low) resolution setting is approximately 4 times higher than when using the Unit/Unit setting, the corresponding noise levels are so high with Low/Low resolution that SNR is dramatically and negatively impacted. In general, the Unit/Unit setting provides the best overall balance of high signal intensity and low noise.
Perhaps even more important than signal and noise considerations, is the impact mass spectral resolution can have on analyte selectivity against co-eluting matrix interferences. When a low resolution setting is used (e.g., 3.0 microns), the m/z range of ion fragments that are allowed to strike the detector is broad, and allows non-specific fragments from compounds other than the target analyte (i.e., matrix interferences) to be included in the measurement. Narrowing the m/z range to Unit (0.8 microns) or High (0.6 microns) resolution minimizes the potential of there being close-eluting contaminants with similar m/z fragments. Compound specificity is achieved by using unique MRM transitions with customized resolution settings for each compound, and provides clean MRM chromatograms even in the presence of matrix interferences with common precursor or product ions.
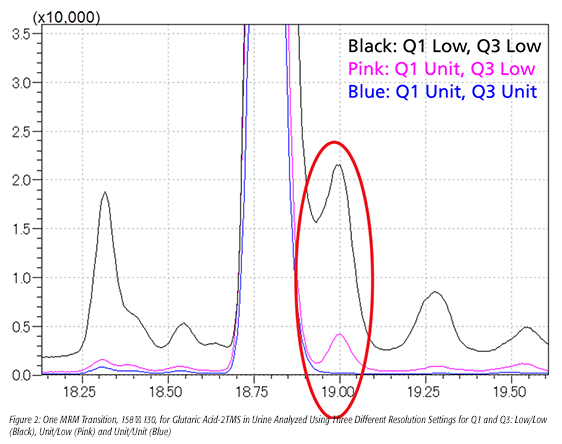 This principle is illustrated in Figure 2. When using Low/Low resolution settings (black trace), the m/z 158 → 130 transition for glutaric acid-2TMS in this urine sample was subject to interference from close-eluting contaminants with similar precursor-product ion pairs. As the resolution was increased, the m/z range of fragments allowed to strike the detector was narrowed from 3.0 to 0.8 microns, and the interference was eliminated. The signal intensity is reduced with Unit/Unit resolution, but this is due primarily to eliminating the fragments from sources other than the target compound, and makes the resulting quantitation of the target more accurate.
This principle is illustrated in Figure 2. When using Low/Low resolution settings (black trace), the m/z 158 → 130 transition for glutaric acid-2TMS in this urine sample was subject to interference from close-eluting contaminants with similar precursor-product ion pairs. As the resolution was increased, the m/z range of fragments allowed to strike the detector was narrowed from 3.0 to 0.8 microns, and the interference was eliminated. The signal intensity is reduced with Unit/Unit resolution, but this is due primarily to eliminating the fragments from sources other than the target compound, and makes the resulting quantitation of the target more accurate.
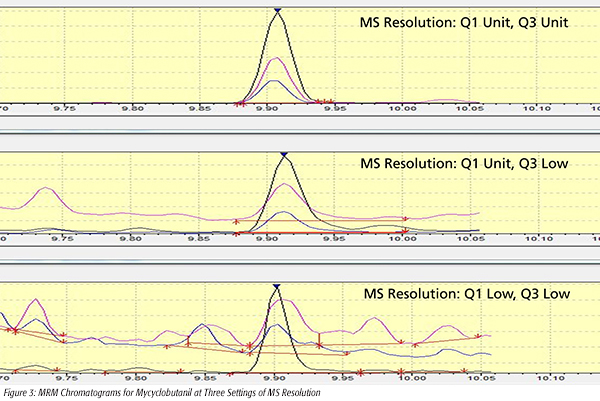 The effect of resolution settings on compound selectivity can also be seen in the matrix-matched pesticide standards used for the baby food project. Figure 3 shows the overlaid MRM chromatograms for mycyclobutanil (1.0 ng/mL) at three different resolution settings for Q1 and Q3: Low/Low, Unit/Low and Unit/Unit. When using the Low/Low resolution setting, background interference from the baby food matrix is clearly evident, and produces interferences which prevent proper integration and confirmation of compound identity using peak ratios. The Unit/Unit resolution setting narrowed the m/z range on both Q1 and Q3 to eliminate the fragments from close-eluting matrix interference, and prevent them from contributing to the signal. The peak was easily and accurately integrated, and peak area ratios for the three individual transitions can be used for confirmation of compound identity.
The effect of resolution settings on compound selectivity can also be seen in the matrix-matched pesticide standards used for the baby food project. Figure 3 shows the overlaid MRM chromatograms for mycyclobutanil (1.0 ng/mL) at three different resolution settings for Q1 and Q3: Low/Low, Unit/Low and Unit/Unit. When using the Low/Low resolution setting, background interference from the baby food matrix is clearly evident, and produces interferences which prevent proper integration and confirmation of compound identity using peak ratios. The Unit/Unit resolution setting narrowed the m/z range on both Q1 and Q3 to eliminate the fragments from close-eluting matrix interference, and prevent them from contributing to the signal. The peak was easily and accurately integrated, and peak area ratios for the three individual transitions can be used for confirmation of compound identity.
For most analyses, the Unit/Unit resolution settings for Q1 and Q3 provide the best combination of sensitivity and selectivity. Lower resolution settings (Unit/Low or Low/Low) allow matrix contaminants to interfere with quantitation, and are not recommended. Higher resolution settings (e.g., High/Unit or High/High) which narrow the m/z range to 0.6 microns on one or both sets of quadrupoles can also be used when matrix interference is severe. In this case, signal intensity will be reduced, but interference will be eliminated and quantitation accuracy will be improved.
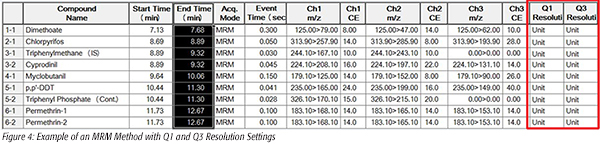 Mass spectral resolution for Q1 and Q3 on the GCMS-TQ8030 can be adjusted individually for each “event,” or set of MRM transitions for a specific analyte. With this feature, mass spectral resolution in an analytical method can easily be customized to optimize response and chromatographic selectivity for each analyte. An example method illustrating this customization is shown in Figure 4.
Mass spectral resolution for Q1 and Q3 on the GCMS-TQ8030 can be adjusted individually for each “event,” or set of MRM transitions for a specific analyte. With this feature, mass spectral resolution in an analytical method can easily be customized to optimize response and chromatographic selectivity for each analyte. An example method illustrating this customization is shown in Figure 4.
Conclusion
Detection of pesticides was demonstrated at single-digit ng/mL (ppb) levels in a complex sample matrix, and linear calibration was confirmed from 1–200 ng/mL in matrix-matched standards. Precision and accuracy were established by replicate analyses of matrix spiked aliquots at 1.0 and 5.0 ng/mL, and found to have single-digit %RSD for those pesticides that did not appear in the sample matrix. Approximately one-third of the pesticides in this study were also detected in the un-spiked baby food extract at concentration below 1.0 ng/mL. The concentration range used in this study covers the maximum residue levels (MRLs) for many pesticides, validating the utility of the MRM mode for analysis of pesticides in complex food matrices.
For most analyses, the Unit/Unit resolution settings for Q1 and Q3 provide the best combination of sensitivity and selectivity. Lower resolution settings (Unit/Low or Low/Low) allow matrix contaminants to interfere with quantitation, and are not recommended. Higher resolution settings (e.g., High/Unit or High/High) which narrow the m/z range to 0.6 microns on one or both sets of quadrupoles can also be used when matrix interference is severe. In this case, signal intensity will be reduced, but interference will be minimized or eliminated, and quantitation accuracy will be improved. A powerful feature of the Shimadzu GCMS-TQ8030 is the ability to adjust the Q1 and Q3 resolution settings individually to customize the method.
A Shimadzu GCMS-TQ8030 system operated in the MRM mode was shown to be a rapid, sensitive, and selective technique for analysis of various classes of pesticides in baby foods in the range required for many regulatory MRLs. Reliable, precise measurements were obtained for 36 pesticides. The Shimadzu GC/MS/MS Pesticide Database simplified development of the MRM method.
The authors wish to acknowledge Restek Corporation, Bellefonte, PA for useful discussions and advice regarding chromatographic conditions and column selection. Restek also provided the GC columns, standards and the QuEChERS sample extracts used in this study.
References
1. AOAC Official Method 2007.01. 2007. Pesticide residues in foods by acetonitrile extraction and partitioning with magnesium sulfate.
2. Shimadzu GC/MS/MS Pesticide Database. 2012.
3. Shimadzu Application News. 2013. Analysis of organophosphorus pesticides in baby foods using a triple-quadrupole GC/MS/MS system. No. GCMS-1304.
For more information, please visit shimadzu.com.
Looking for quick answers on food safety topics?
Try Ask FSM, our new smart AI search tool.
Ask FSM →





